
RES E AR C H Open Access
A three-dimensional collagen construct to model
lipopolysaccharide-induced activation of BV2
microglia
Randy Tatt Yhew Haw
1†
, Chih Kong Tong
1†
, Andrew Yew
1
, Han Chung Lee
2
, James B Phillips
3
and Sharmili Vidyadaran
1*
Abstract
Background: We report a novel method of culturing microglia in three dimension (3D) using collagen as a
substrate. By culturing microglia within a matrix, we aim to emulate the physical state of microglia embedded
within parenchyma.
Methods: BV2 microglia cell suspensions were prepared with type I collagen and cast into culture plates. To
characterise the BV2 microglia cultured in 3D, the cultures were evaluated for their viability, cell morphology and
response to lipopolysaccharide (LPS) activation. Conventional monolayer cultures (grown on uncoated and
collagen-coated polystyrene) were set up concurrently for compa rison.
Results: BV2 microglia in 3D collagen matrices were viable at 48 hrs of culture and exhibit a ramified morphology
with multiplanar cytoplasmic projections. Following stimulation with 1 μg/ml LPS, microglia cultured in 3D collagen
gels increase their expression of nitric oxide (NO) and CD40, indicating their capac ity to become activated within
the matrix. Up to 97.8% of BV2 microglia grown in 3D cultures gained CD40 positivity in response to LPS, compared
to approximately 60% of cells grown in a monolayer (P < .05). BV2 microglia in 3D collagen gels also showed
increased mRNA and protein expression of inflammatory cytokines IL-6, TNF-α and the chemoattractant MCP-1
following LPS stimulation.
Conclusions: In summary, BV2 microglia cultured in 3D collagen hydrogels exhibit multiplanar cytoplasmic
projections and undergo a characteristic and robust activation response to LPS. This culture system is accessible to
a wide range of analyses and provides a useful new in vitro tool for research into microglial activation.
Keywords: Microglia, Lipopolysaccharide, Collagen matrix, Three-dimensional cultures
Background
Microglia are tissue-specific macrophages of the central
ner vous system (CNS) and derive from primitive haem-
atopoietic progenitors of erythromyeloid origin [1,2].
These mononuclear phagocytes are the resident im-
mune cells of the CN S, along with other subsets of
mononuclear phagocytes including meningeal macro-
phages, choroid plexus macrophages and perivascular
macrophages [3]. Microglia in the brain are disseminated
throughout the parenchyma and are highly motile. In the
healthy mature CNS, microglia exist mainly in a ramified
form, continuously traversing the CNS and using their cyto-
plasmic processes to sample the tissue environment [4].
Although microglia have long been thought of as merely
a stromal cell, microglia research has now revealed crucial
roles for these cells in inflammation and in different stages
of neurodevelopment. Homeostatic changes in the CNS
rapidly trigger a reactive form of microglia, characterised
by a shift to amoeboidal morphology, increased motility,
proliferation and release of inflammatory mediators [5].
In the embryonic brain, microglia associate closely with
apoptotic cells, presumably to promote developmental
neuron death and phagocytose the ensuing cellular debris
†
Equal contributors
1
Neuroinflammation Group, Immunology Laboratory, Department of
Pathology, Faculty of Medicine & Health Sciences, Universiti Putra Malaysia,
43400 Serdang, Selangor, Malaysia
Full list of author information is available at the end of the article
JOURNAL OF
NEUROINFLAMMATION
© 2014 Haw et al.; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative
Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and
reproduction in any medium, provided the original work is properly credited. The Creative Commons Public Domain
Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article,
unless otherwise stated.
Haw et al. Journal of Neuroinflammation 2014, 11:134
http://www.jneuroinflammation.com/content/11/1/134
[6]. During postnatal development, microglia play a role
in synaptic pruning by engulfing synaptic material [7,8]. In
adult hippocampal neurogenesis, microglia provide an
important housekeeping role by phagocytosing and clear-
ing apopto tic ne wborn neurons [9]. These function s of
the microglia have led to the recognition of the signifi-
cance of this cell in brain rese arch.
Homogenous cell cultures are a valuable approach for
neuroscience research that allows monitoring of the cell
population of interest in a carefully controlled environ-
ment. Monolayer cultures fail to recreate the 3D spatial
arrangement of cells and matrices present in tissues, and
stiff plastic substrates do not resemble the physical environ-
ment of the CNS. Culture systems that better mimic the
behaviour of microglia within a 3D milieu would allow
scrutiny of these cells in a more relevant tissue-like micro-
environment. Brain slice culture allows in situ examination
of cellular responses, and has been used to study microglial
responses to carcinomas [10] and cerebral amyloidosis [11].
However, downstream analysis of individual cell popula-
tions can be complicated and the local cellular environment
is complex.
Here, we describe a 3D culture system utilising type I
collagen as the basal substrate to provide a matrix for the
culture of microglial cells. Type I collagen is a matrix
material that is easily manipulated, is widely used in
culture models [12], and has previously been used to
develop 3D cultures for astrocytes and neurons [13-15].
Being a simple matrix, it also serves as a suitable baseline
scaffold on which the deposition of other extracellular
matrix (ECM) molecules can be detected. With control
over the seeding density and the chemical environment
within the gels, we are able to examine specific features of
microglia in a 3D matrix with ease of monitoring the cells
compared to in vivo models or brain slice cultures.
In the laboratory we routinely culture BV2 microglia, a cell
line of murine origin immortalised with v-raf/v-myc onco-
genes and commonly used in microglia studies. The BV2
microglia are similar in morphology to isolated microglia,
express inflammatory mediators and display phagocytic
activity [16]. To stimulate the BV2 microglia into an inflam-
matory phenotype, the bacterial cell wall component lipo-
polysaccharide (LPS) is used. By examining morphology,
viability and activation status (by evaluating nitric oxide
production along with CD40 and inflammatory cytokine
expression) of BV2 microglia in 3D constructs and com-
paring them to conventional monolayer cultures, we char-
acterise microglia cultured in 3D and report a model for
microglial activation in a 3D collagen matrix using LPS.
Methods
BV2 cells
BV2, an immortalised mouse microglia cell line, was cul-
tured in high glucose Dulbecco modified Eagle medium
(DMEM; Gibco, Carls bad USA) supplemented with 5%
foetal bovine serum (Gibco, Carlsbad USA), 6.25 μg/ml
insulin (Sigma-Aldrich, St. Louis USA), 1X non essen-
tial amino acid (Gib co, Carlsbad USA), 1% penicillin
and streptomycin ( i-DNA , Singapore), 0.5% fungizone
(Gibco, Carlsbad USA) and 0.1% gentamicin (Gib co,
Carlsbad USA). The cultures were maintained in a hu-
midified incubator at 37°C with 5% CO
2
:95% air.
Culture of BV2 cells on monolayer surfaces and in
three-dimensional collagen gels
BV2 cells were har vested at 70-90% confluency and
counted u sing a haemocytometer. Cultures up to pas-
sage 20 were used for experiments. The seeding density
for all culture conditions and downstream assays was
0.3 × 10
6
cells/well. For uncoated and collagen-coated
monolayer cultures, cells were seeded in 6-well plates.
For coated monolayer cultures, culture plates were pre -
coated with type I rat tail collagen (First Link UK Ltd,
Birmingham, UK) in 0.6% acetic acid for 30 min. The
culture flasks were then rinsed with supplemented DMEM
to remove any traces of acid.
For 3D cultures, collagen gels were prepared by adding
10% v/v cell suspension in supplemented DMEM, 10% v/v
10X minimum essential medium (MEM; Sigma-Aldrich St.
Louis USA) and 80% v/v type I rat tail collagen (2 mg/ml in
0.6% acetic acid; First Link, Birmingham UK). The MEM
and collagen were mixed and neutralised using sodium
hydroxide as assessed by colour change of the phenol red
indicator. Upon neutralisation, the collagen-MEM mixture
was gently mixed with the BV2 cell suspension and
transferred to culture plates (0.3 ml/well in 24-well
plates and 1.2 ml/well in 6-well plates or 35 mm WillCo
(WillCo Wells, Amsterdam, Netherlands) dishes; resulting
gels were approximately 2-mm thick). The gels were
allowed to set in a humidified incubator at 37°C with 5%
CO
2
:95% air for 5 to 10 min and subsequently covered
with 2 ml supplemented DMEM. The cultures were main-
tained for up to 48 hrs and cells were retrieved for analysis
using 0.25% trypsin for monolayer cultures and 0.125%
type I collagenase for 3D cultures.
To stimulate the cultures, BV2 microglia were treated
with 1 μg/ml lipopolysaccharide (LPS; E. coli serotype
O26:B6; Sigma-Aldrich St. Louis USA, Cat. No. L2762) in
supplemented DMEM. Controls were subjected to media
change only.
Scanning electron microscopy
Collagen gels with and without BV2 cells were processed
based on a mo dified version of Lizárrag a and col-
leagues [17]. Briefly, samples were p refixed in 2.5% v/v
glutaraldehyde at 4°C for 4 hrs and washed with 0.1 M
sodium cacodylate buffer. After overnight incubation,
samples were post-fixated in 1% osmium tetraoxide.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 2 of 10
http://www.jneuroinflammation.com/content/11/1/134
Samples were then dehydrated using a graded series of
ethanol followed by immersion in 100% acetone. The
samples were then transferred to a Baltec CPD 030 Critical
Point Dryer for critical point drying. Samples were coated
with gold-palladium in a Baltec SCD 005 Sputter Coater
and examined under JEOL JSM-6400 SEM.
DAPI/propidium iodide staining
BV2 cell viability in all culture conditions was assessed
using propidium iodide ( PI) and a 4',6-d iamidino-2-
phenylindole, dihydrochlorid e (DAPI) counterstain. In
brief, 20 μg/ml PI (Molecular Probes, Oregon USA)
wasaddedtoculturesat24and48hrsandincubated
for 10 min at 37°C. Supernatant was t hen removed and
the cultures rinsed in 1× PBS thrice for 5 min to remove
PI residue. Cells were fixed with 4% paraformaldehyde
(PFA) at 4°C for 1 hr. The cultures were then incubated
with 1 μg/ml DAPI (Molecular Probes, Oregon USA) in
1X PBS with 0.1% Triton-X for 10 min. For the positive
control, Triton-X (0.2% in basic DMEM) was used to treat
the cells for 5 min prior to PI staining.
Lactate dehydrogenase assay
Lactate dehydrogenase (LDH) assay is a colourimetric assay
that specifically detects the enzyme lactate dehydrogenase.
This enzyme is particularly stable and is present in the cul-
ture supernatant when cells are damaged. For this assay,
three controls were established, namely, the background
control (only media), low control (untreated cells) and posi-
tive control (cells treated with lysis solution). All cultures
were supplied with an equal volume (1.5 ml) of culture
medium. Following overnight incubation at 37°C and 5%
CO
2
:95% air, cells were treated with 1 μg/ml LPS in phenol
red-free, supplemented DMEM or subjected to a media
change for untreated cells. At 48 hrs, lysis solution obtained
from the Cytotoxicity LDH kit (Roche, Mannheim Germany)
was added into the positive control wells. Next, 500 μlof
media from each well was aliquoted into tubes and centri-
fuged at 1,500 rpm for 5 min to remove any debris. Next,
100 μl of the supernatant was transferred into a 96-well
plate in triplicate. A reaction mixture was prepared by
mixing the catalyst solution and dye solution from the kit
and 100 μl of the reaction mixture was added into each
well. The 96-well plate was incubated for 20 min prior to
measuring the absorbance at a 490-nm wavelength using
a microplate reader (Dynex, Virginia United States). Read-
ings from background controls were subtracted from all
reaction absorbance readings.
Lectin staining
BV2 cell morphology was assessed using lectin histochem-
ical staining. Briefly, cells were fixed with 4% paraformalde-
hyde (PFA) in 1X PBS for 1 hr followed by permeabilisation
with 0.2% Triton- X in 1× PBS for 30 min. Cells were then
incubated with fluorescein isothiocyanate (FIT C)-conjugated
tomato lectin (1:300 dilution in 0.2% of Triton- X in 1X PBS;
Sigma-Aldrich, St Louis USA) for 1 hr. Nuclei were counter-
stained with 0.1 μg/ml DAPI for 5 min before cells were
viewed and photographed with an inverted fluorescence
microscope (Olympus, Tokyo Japan) and laser scanning
confocal microscope (Leica DMIL, Wetzlar Germany).
CD40 immunophenotyping
For immunopheno typing, cells were harvested from cul-
tures and res uspended in 100 μl of 1× PBS. They were
then incubated with Fixable Viability Dye eFluor™ 780
(eBioscience, San Diego U SA) at 1:1000 dilution for
15 min, followed by an anti-mouse CD40-FIT C antibody
(1:100 dilution; BD Pharmigen, San Diego USA) for 30 min
at 4°C. Cells were then washed and resuspended in 1× PBS
before being analysed by a FACS Fortessa Cytometer
(BD Biosciences, San Jose, CA USA). Gating was applied
to identify intact cells according to forward a nd side
scatter plots; dead cells were excluded from analysis using
the Fixable Viability Dye eFluor™ 780 staining. Data were
analysed using the FACS Diva software.
Griess assay
This assay was performed to study the activation of BV2
cells in a resting state and LPS-treated state by determining
the production of nitric oxide (NO) at 36 and 48 hrs, re-
spectively. BV2 cells were seeded in 6-well plates for mono-
layer cultures and 24-well plates for 3D collagen cultures at
a seeding density of 0.3 × 10
6
cells/well. All cultures were
supplied with an equal volume (2 ml) of culture medium.
Following overnight incubation at 37°C and 5% CO
2
,cells
were treated with 1 μg/ml LPS in phenol red-free, supple-
mented DMEM. For the Griess assay, 200 μlofmediafrom
each well was aliquoted into tubes and centrifuged at
1,500 rpm for 5 min to remove any debris. Next, 50 μlof
the supernatant was transferred into a 96-well plate in trip-
licate. A series of sodium nitrite (NaNO
2
)standardsforthe
Griess assay were prepared by serial dilution, ranging from
0 μMto100μM. Griess reagent was freshly prepared by
dissolving sulphanilamide (Sigma-Aldrich, United States)
and N-(1Naphtyl)ethylenediamine (Sigma-Aldrich, St Louis
United States) with phosphoric acid. Fifty microliters of
Griess reagent was added to each well before absor-
bances were measured with a microplate s pec trometer
(Dynex, Virginia United States) at 530 nm wavelength.
The NO
2
−
concentr ation was then evaluated by normalising
the absorbance reading using the graph equation obtained
from the plotted standard graph.
Detection of cytokine expression with reverse transcriptase
quantitative PCR and a cytokine bead array
The mRNA expression of MCP-1, IL-6, IL-1β,IL-12β
and TNF-α of BV2 mic roglia was assessed with reverse-
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 3 of 10
http://www.jneuroinflammation.com/content/11/1/134

transcriptase quantitative P CR (RT-qP CR). The RNA
of BV2 microglia wa s isolated using the RNeas y Plus
Mini Kit (Qiagen, Limburg German y) after a 6-hr
stimulation with LPS. The isolatio n process wa s con-
ducted according to the kit’s manual. The yield of
total RNA was quantified by optical density (OD)
readings at 260 nm, and the purity wa s estimated by
the 260:280 nm ratio. Using the SuperScript III
Reverse Transcriptase Kit (Invitrogen, Carlsbad U SA),
an equivalent amount of RNA samples (500 ng) were
then re verse transcribed into single stranded cDNA in
a reaction mixture consisting of 0.5 mM dN TP mix,
2.5uM olig o d( T)
20
,0.01MDTT,40URNAseOUT™,
1× RT buffer and 200U Superscript™ RT prepared in
20uL per reaction, according to the manufacturer’s
protocol. The primers and housekeeping genes (Hmbs,
Pgk1 and Psmb2) were designed and probes selec ted
using ProbeFinder Version 2.49 (Universal Probe Library
Assay (UPL) Design Center, Roche). Details on the primer
sequences are in Additional file 1. PCR wa s then per-
formed on a LightCycler 480 System (Roche, B a sel
Switzerland).Conditions for the RT-qPCR were a pre-
denaturing step of 95°C for 10 min; 45 cycles of 95°C
for 10 sec , 60°C for 30 se c, and 72°C for 1 sec; and
finishing with a cooling step at 40°C for 30 sec [18].
A PCR efficiency of between 90 and 110% and an
R-squared value >0.98 were used to define successful
assays. Relative quantification of target g ene e xpression
in our samples wa s carried out using the comparative
Ct method. We performed intra-sample data normalisa-
tion against the three endogenous control reference
genesasmentionedabove.
For the bead array, culture supernatants were assesse d
at 48 hrs for expression of IL-6, IL-10, MCP-1, IFN-γ,
TNF, and IL-12p70 using the multiplex bead array kit
(BD Cytometric Bead Array mouse inflammation kit,
BD Biosciences, San Jose, C A, USA). The samples were
assayed according to the manufacturer ’s instructions
with the FACS Fortessa flow cytometer (BD Bioscienc e,
San Jose, CA, USA). The resulting data were analysed
with FC AP array software (BD Bioscience, San Jose, CA,
USA). Expression of the cytokines wa s e valuated by
determining their respe ctive concentration (pictograms
per millilitre) using individual standard curves.
Statistical analysis
Signific ance was assessed using GraphPad Prism version 6
(GraphPad Software, CA, USA, http://www.graphpad.com/).
Results
Scanning electron microscopy of BV2 microglia and
collagen gel structure
Examination of the collagen gel structure was performed
using scanning electron microscopy. The matrix consists
of a dense fibrillar network, with numerous interfibrillar
spaces (Figure 1A). Microglia were detected on the surface
of the gels and also beneath the surface (arrows and
arrowheads in Figure 1B,C ).
Morphology and viability of BV2 microglia cultured in
monolayer and three dimensions
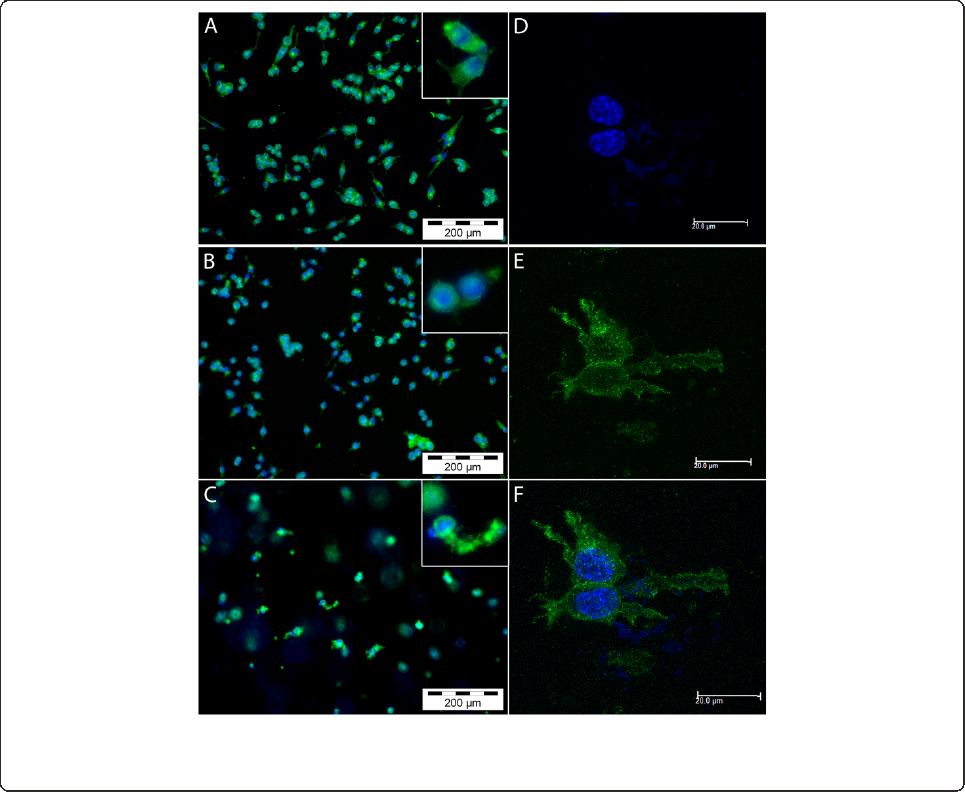
To examine the morphology of microglia in the various
culture systems , we performed staining of cells with
FITC-tagged lectin. M icroglia cultured on uncoated
monolayer surfaces mostly exhibited round cytopla sm
with some bipolar project ions (Figure 2A). In collagen-
coated monolayer cultures , the extent of amoeboidal
morphology seemed increased (Figure 2B) with cyto-
plasmic area appearing minimal, indicating the extent
of deramification. The morphology of cells cultured in
3D collagen wa s distinct from monolayer, with clear
multiplanar projections (Figure 2C). The microglia
were suspended within the collagen matrix and ev enly
distributed across the width of the gel. When viewed
with confocal microscopy, the extent of ramification of
microglia cultured in the 3D matrix was evident, with
cells displaying long and multi directional cellular pro-
jections (Figure 2F and video in Additional file 2).
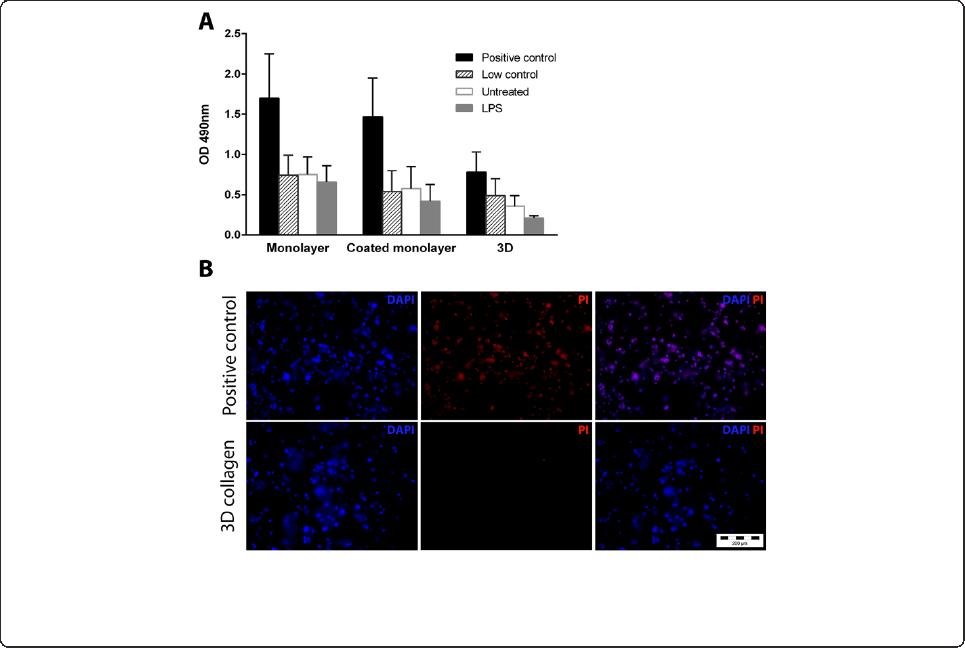
To determine whether microglia cultured in 3D were
viable, the lactate dehydrogenase (LDH) assay and DAPI/PI
staining were performed. Compared to the respective posi-
tive controls, BV2 microglia in all three culture formats
Figure 1 Scanning electron micrographs of three-dimensional (3D) collagen gel and microglia cultured in 3D. (A) Structure and
organisation of fibrils of 2 mg/ml collagen gels. (B,C) Low magnification images show microglia on the surface (arrows) and embedded
(arrowheads) within the collagen gels. Magnification as indicated on each micrograph.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 4 of 10
http://www.jneuroinflammation.com/content/11/1/134
showed low LDH release (Figure 3A). Ev en following treat -
ment with 1 μg/ml LPS, LDH levels in the culture superna-
tants remained low. Additionally, BV2 microglia showed
negligible PI staining at 24 (data not sh own) and 48 hrs
post-culture in 3D colla gen (Figure 3B).
CD40 expression of microglia cultured in the
three-dimensional c ollagen matrix
We routinely assess expression of the co-stimulatory
molecule CD40 as a measure of the activation status for
microglia. Upon exposure to lipopolysaccharide (LPS),
microglia secrete inflammatory mediators and upregulate
expression of major histocompatibility complex (MHC)
class II receptors and CD40 to facilitate antigen presenta-
tion to T lymphocytes [19]. Due to these effects, LPS is a
common stimulus for activating microglia in vitro [20-22],
including for our previous work [23-25].
The number of microglia expressing basal CD40 in 3D
cultures was higher (27.6 ± 14.6%) compared to cells
grown in uncoated monolayer cultures (5.1 ± 3.39%)
(P < .05; Kruskal-Wallis with Dunn’smultiplecom-
parisontest).WhenstimulatedwithLPS,thenumber
of CD40
+
BV2 microglia increased by 70% (P < .001),
with almost the entire population of BV2 cells cul-
tured in 3D collagen shifting to a CD40
+
phenotype at
24 hrs (97.8 ± 1.5%; Figure 4A). CD40
+
BV2 cells in
uncoated and collagen-coated monolayer cultures also in-
creased from 5.1 ± 3.4% and 12.8 ± 11.2% to 60.2 ± 20.0%
and 62.3 ± 11.8%, respectively (Figure 4A; P < .05). The
increase in CD40
+
cells was sig nificantly higher f or 3D
cultures compared to uncoated and collagen-coated
monolayer cultures (P < .05; Kruskal-Wallis with Dunn’s
multiple comparison test). Achie ving a hom ogenous
population shift may indicate a uniform le vel of activa-
tion amongst the microglia , an effect that appears to
be rendered by the 3D collagen culture.
The median fluorescence intensity (MFI) readouts on
a flow cytometer indicate whether cells within a positive
population express the marker in question at different
intensities. Using MFI, we show that not only do the
Figure 2 Morphology of microglia cultured in monolayer and three-dimensional (3D) systems. Microglia cultured in monolayer (A), coated
monolayer (B) and 3D collagen gels (C,D,E,F) were stained with fluorescein isothiocyanate (FITC)-tagged lectin and 4',6-diamidino-2-phenylindole,
dihydrochloride (DAPI) and viewed with fluorescent (A-C) or confocal (D-F) microscopy.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 5 of 10
http://www.jneuroinflammation.com/content/11/1/134
number of CD40
+
BV2 microglia increase in 3D culture,
but the degree of their CD40 expression also increases
3-fold compared with cells grown on uncoated mono-
layer and collagen-coated monolayer surfaces (P < .05;
Figure 4B). Also, microglia in monolayer cultures appear
to respond to LPS by acquiring CD40 expression, and
not by increasing the level of expression (Figure 4B).
Collectively, microglia in 3D culture appear to be acti-
vated in a more homogenous fashion, and to a greater
extent than cells in monolayer cultures. Interestingly,
the intensity of CD40 expression in unstimulated cul-
tures remained similar between all three culture formats.
Therefore, although the number of CD40
+
cells is higher
in 3D cultures compared to monolayer, the degree of
expression per cell is similar.
Nitric oxide and inflammatory cytokine expression by
microglia cultured in three dimensions
Disease o r damage within the CNS triggers production
of nitric oxide (NO) by microglia and a strocytes [26].
The inducible nitric oxide synt hase (iNOS) iso form
of NO synthase is responsible for secretion of large,
continuous amounts o f NO by these glial cells during
inflammation, as compared to neuronal NOS (nNOS)
that is constitutively expressed in neurons and is
believed to have a physiological role in the brain [26].
To assess NO production by BV2 microglia in mono-
layer and 3D cultures, cells were seeded in the different
culture formats with equal seeding number per well.
Microglia in monolayer and 3D cultures had negligible
NO production (<3 μM at both 36 and 48 hrs; Figure 5),
indicating that the culture conditions alone do not in-
duce NO expres sion. Following stimulation with LPS,
BV2 microglia in all culture formats showed an induc-
tion of NO expression, approximately 4 to 8 times higher
at 36 hrs and 6 to 10 times higher at 48 hrs compared to
untreated BV2 (P < .05). Negligible NO induction was
detected at 24 hrs post-LPS (data not shown). At 36 hrs,
BV2 grown as uncoated monolayer cultures produced
the highest amount of NO (20.03 ± 0.76 μM), followed
by collagen-coated monolayer cultures (6.59 ± 2.39 μM)
and 3D cultures (10.54 ± 1.46 μM). These levels in-
creased for uncoated monolayer, collagen-coated mono-
layer and 3D culture a t the 48-hr time point to 26.96 ±
1.76 μM, 17.87 ± 2.27 μM and 24.47 ± 1.75 μMNO,
respectively. The NO expression by LPS-treated BV2
cells in 3D cultures was not significantly different com-
pared to microglia cultured in uncoated or collagen-
Figure 3 BV2 microglia are viable in three-dimensional (3D) collagen gels. (A) BV2 microglia in monolayer, coated monolayer and 3D
culture conditions were assessed for lactate dehydrogenase (LDH) activity at 48 hrs. Data are mean ± SD from three independent experiments.
(B) Cells were stained with 4',6-diamidino-2-phenylindole, dihydrochloride (DAPI) and propidium iodide (PI) to assess viability at 48 hrs
post-culture. LPS, lipopolysaccharide.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 6 of 10
http://www.jneuroinflammation.com/content/11/1/134
coated monolayer cultures at either time points (P >.05;
Kruskal-Wallis with Dunn’s multiple comparison test).
The response of microglia in 3D cultures to LPS was
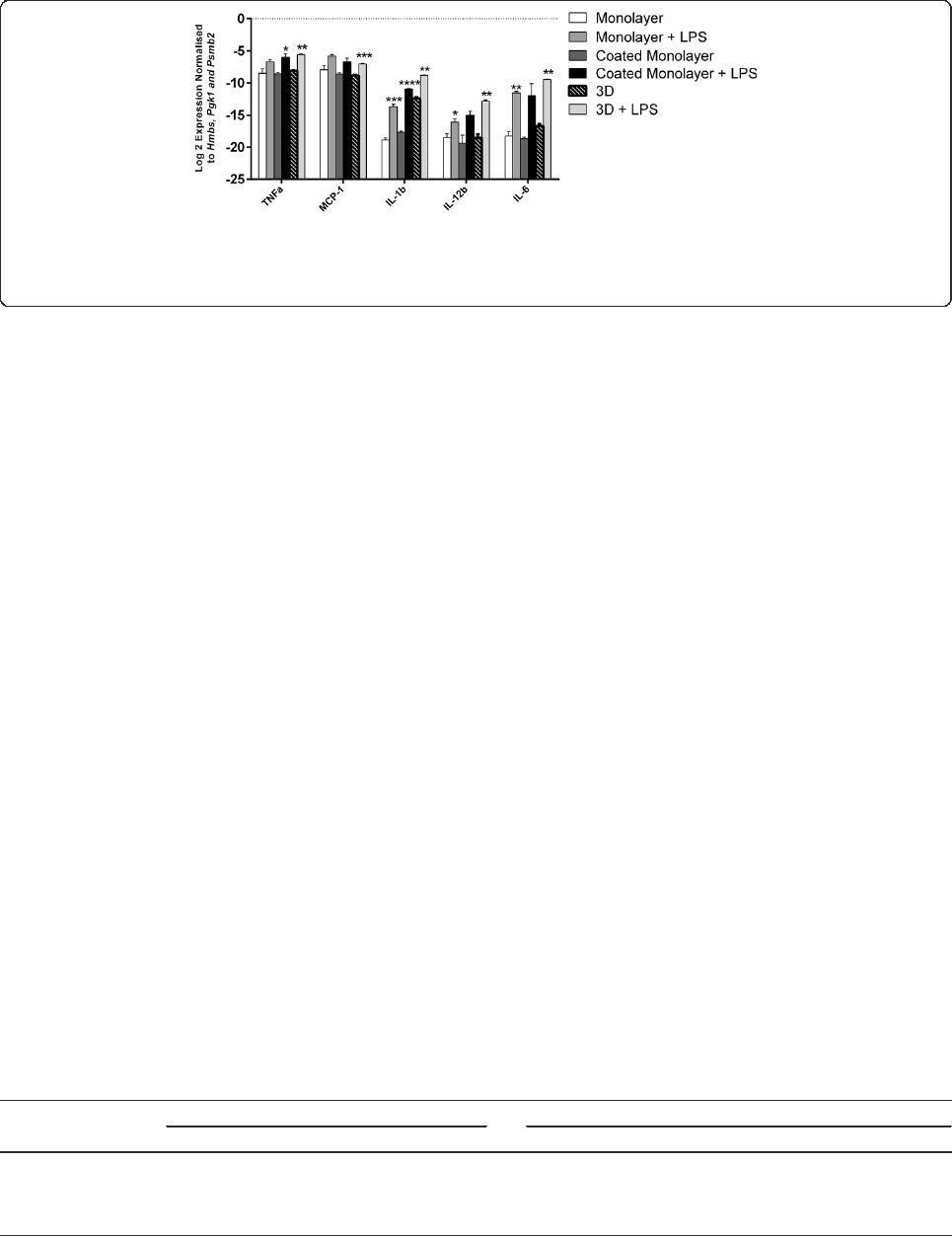
also evaluated by determining expression of inflammatory
cytokines mRNA and protein. At 6 hrs, mRNA expression
of all inflammatory cytokines tested (MCP-1, IL-6, IL-1β,
IL-12β and TNF-α) was significantly upregulated in BV2
microglia cultured in 3D, relative to housekeeping gene
expression (Figure 6 and Additional file 3). Similar to
mRNA expression, protein levels for IL-6, TNF-α and
MCP-1 were increased following LPS stimulation in 3D
cultures. At 48 hrs of LPS stimulation, BV2 microglia in
3D cultures re corded signif icantly higher expression of
IL-6, TNF-α and MCP-1 compared to untreated BV2
cells (P < .05; Mann Whitney Test) (Table 1). Levels of
IL-6, TNF-α and MCP-1 increa sed by 1 ,998.3 pg/ml,
1,735.9 pg/ml and 5,119.0 pg/ml res pe ctively compared
to basal (untreated) levels. As we have shown before [23],
levels of IFN-γ,IL-10and IL-12p70 were unaffected by
LPS stimulation. Additionally it was observed that un-
stimulated BV2 microglia in 3D cultures demonstrated
markedly lower MCP-1 levels compared to monolayer
cultures (P < .05; Kruskal-Wallis with Dunn’s multiple
comparison test), which was significantly upregulated with
LPS stimulation. MCP-1 expression for LPS-treated BV2
cells in monolayer and coated monolayer cultures was not
significantly different from unstimulated cultures.
Discussion
Conventional monolayer cell cultures involve the growth
of cells on a plastic surface, often coated with extracellu-
lar matrix proteins such as collagen and laminin to
encourage adherence. Several fundamental disparities
exist between these conventional monolayer cultures
and cells in situ, namely cells cultured in monolayer
grow flat, may receive cues from the stiff matrix, do not
grow in a stratified manner and have only one side
adhering to the plastic surface. This also means that
perfusion of nutrients for the cells only occurs via the
non-adhered side. For highly reactive cells such as the
microglia, these culture conditions could affect their
behaviour. The BV2 microglia cell line is commonly
used for microglia research, is well-characterised and is
routinely studied in our laboratory. To approach in vitro
BV2 microglia cultures in a more relevant manner, we
sought to culture BV2 microglia within a collagen matrix
to mimic the mechanical relationship of these cells with
tissue. We demonstrate that BV2 microglia grown in 3D
collagen are viable, embedded and distributed within
the matrix, with a ramified morphology. The ability of
microglia to grow within collagen matrices also allows
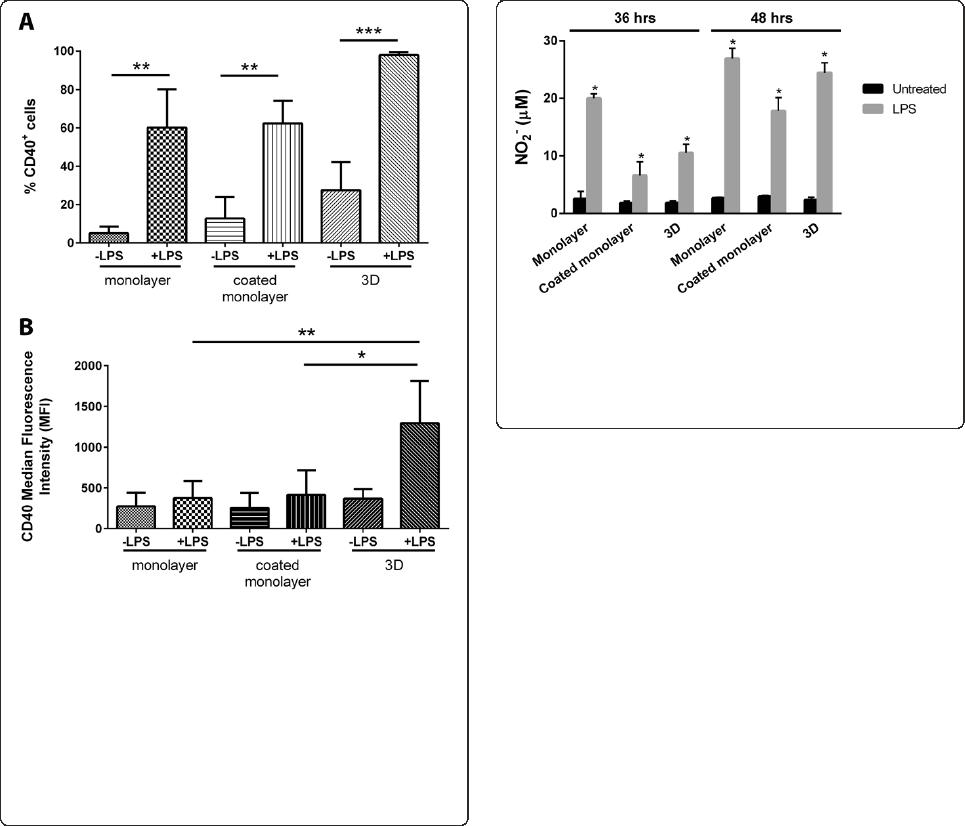
Figure 4 BV2 microglia in three-dimensional (3D) collagen gels
shift as a population to express CD40 in response to lipopoly-
saccharide (LPS) stimulation. BV2 microglia in monolayer, coated
monolayer and 3D culture conditions were activated with 1 μg/ml
LPS (+LPS) and CD40 expression analysed with flow cytometry at
24 hrs. (A) Histograms show percentage of CD40
+
cells. Data are
mean ± SD from three independent experiments. **P < .01, ****P
< .001; one-tailed Mann-Whitney U test. (B) Histograms show median
fluorescence intensity (MFI) of CD40
+
. Data are mean ± SD from three
independent experiments. *P <.05,**P < .01 ; Kruskal Wallis with Dunn’s
Multiple Comparison test.
Figure 5 BV2 microglia cultured in three-dimensional (3D)
collagen gels express nitric oxide (NO) in response to
lipopolysaccharide (LPS) stimulation. BV2 microglia in monolayer,
coated monolayer and 3D culture conditions were activated with
1 μg/ml LPS and assessed for NO
2
−
expression at 36 and 48 hrs with the
Griess assay. Data are mean ± SD from three independent experiments.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 7 of 10
http://www.jneuroinflammation.com/content/11/1/134

for multiplanar projections of the cytoplasm, which
would be impossible to achieve with conventional mono-
layer cultures.
Using lipopolysaccharide (LPS), we developed an acti-
vation model for BV2 microglia cultured in 3D. LPS is a
bacterial cell wall component that triggers microglia
activation via Toll-like receptor 4 [19]. With 1 μg/ml
LPS, a dose ro utinely used to activate micr oglia in con-
ventional monolayer cultures [20,24,25], BV2 microglia
in 3D cultures significantly increa sed expression of the
cell surface re ceptor CD40, nitric oxide (NO), IL-6,
TNF-α and MC P - 1. This demonstrates that both cells
and supernatant of 3D collagen cultures can be assayed,
demonstrating the suitability of this cul ture system to
accommodate a range of tests. For CD40 expression,
microglia cultured in 3D show ed a higher number of
cells with ba sal e xpression of CD40 compared to the
monolayer cultures. The NO levels among a ll three
cultures in unstimulated conditions, however, were
similar, indicating that collagen alone (in monolayer
or 3D format s) does not trigger NO product ion in
unstimulated BV2 cells. The extent of CD40 expression
per cell (shown by MFI readings) was also similar
between microglia cultured in monolayer and 3D.
Importantly, LPS stimulation triggers a uniform upreg-
ulation of CD40 in 3D cultures compared to cells
cultured in monolayer formats, with the entire population
of microglia acquiring CD40 expression. Therefore, it
appears that BV2 microglia cultured in 3 D acquire
CD40 towards LPS in a more homogenous manner. The
microglia cultured in 3D also expressed significant
amounts of NO in response to LPS.
Beyond the parameters assayed here, the 3D collagen
culture model may be appropriate for studying micro-
glial deposition of ECM material as it offers a matrix of
simple composition compared to other more complex
substrates such as Matrigel™ [14]. With the data now to
show that BV2 microglia are activated by LPS within 3D
collagen cultures, we are keen to utilise this model for
our main research approach of modulating inflammatory
responses of microglia. The growth of microglia within
an environment that is more physically and spatially
relevant than conventional flat plastic culture plates,
along with the ability to stimulate the microglia into an
activated phenotype, gives us access to a more refined
in vitro tool for microglia research.
Conclusions
By cultur ing micro glia w ithin a simple matrix , we offer
amorerelevantin v itro model compared to conven-
tional monolayer cultures where microglia grow flat on
a plastic surface. BV2 micro glia cultured in 3D co lla-
gen constructs render cells that are viable, well distrib-
uted within the matrix, ramified in morphology and
activated into an inflammatory phenotype following
LPS stimulation.
Figure 6 Reverse-transcriptase quantitative PCR ( RT-qPCR) demonstrates the level of inflammatory cytokines by BV2 microglia 6 hrs
after lipopolysaccharide (LPS) stimulation. The values have been normalised to three housekeeping genes (Hmbs, Pgk1 and Psmb2). TNF,
MCP-1, IL-1b, IL-6 and IL-12b were upregulated in stimulated BV2 microglia for three-dimensional (3D) cultures. Data are mean ± SEM from three
independent experiments. *P < .05, **P < .01, ***P < .001, ****P < .0001; Welch’s t test.
Table 1 BV2 microglia in three-dimensional (3D) cultures show increased expression of IL-6, TNF-α and MCP-1 at
48 hours post-lipopolysaccharide (LPS) stimulation
Untreated LPS-treated
IL-6 TNF-α MCP-1 IL-6 TNF-α MCP-1
Monolayer 1.7 ± 0.3 252.3 ± 120.5 9 447.0 ± 226.8 2 476.0 ± 833.6** 3 823.0 ± 1189.0** 10 459.0 ± 1, 639.0
ns
Coated monolayer 1.6 ± 0.99 217.6 ± 181.8 9 231.0 ± 2268.0 1 764.0 ± 324.3** 2 357.0 ± 500.9** 10 936.0 ± 1292.0
ns
3D 0.7 ± 0.8 8.1 ± 3.1 284.0 ± 73.5 1 999.0 ± 685.2* 1 744.0 ± 911.6* 5 403.0 ± 517.6*
Values are expressed in pg/ml, and assayed using the Cytometric Bead Array. Data are mean ± SD from five independent experiments. *p < .05, **p < .01;
two-tailed Mann-Whitney U test, as compared to untreated controls. ns, not significant.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 8 of 10
http://www.jneuroinflammation.com/content/11/1/134

Additional files
Additional file 1: List of primers and UPL probes used for
reverse-transcriptase quantitative PCR (RT-qPCR) validations.
Additional file 2: Video demonstrating three-dimensional (3D)
image of a BV2 microglia with the 3D collagen gel.
Additional file 3: Reverse-transcriptase quantitative PCR (RT-qPCR)
analyses of TNF, MCP-1, IL-b, IL-12b and IL-6 mRNA.
Abbreviations
CNS: Central nervous system; DAPI: 4',6-diamidino-2-phenylindole,
dihydrochloride; ECM: Extracellular matrix; FITC: Fluorescein isothiocyanate;
iNOS: inducible nitric oxide synthase; LDH: Lactate dehydrogenase;
LPS: Lipopolysaccha ride; MFI: Median fluorescence intensity; nNOS: neuronal
nitric oxide synthase; NO: Nitric oxide; OD: Optical density;
PFA: Paraformaldehyde; PI: Propidium iodide; 3D: three-dimensional.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
RHTY performed the cell cultures, LDH assay, Griess assay and cytokine bead
array. TCK established the 3D collagen cultures, performed the confocal
microscopy, immunophenotyping and viability staining. AY performed the
scanning electron microscopy. LHC performed the RT-qPCR experiments. SV
and JBP conceptualised the study. SV wrote the manuscript. All authors
analysed the data and read and approved the final manuscript.
Acknowledgements
The authors would like to thank Hi-Tech Instruments Sdn. Bhd. and the Brain
Research Institute Monash Unive rsity Sunway for use of their confocal
microscope, staff of the Electron Micro scopy Unit, Universiti Putra Malaysia
for assistance with the electron microscopy, and Dr. Michael Ling King Hwa
for his expert advice on RT-qPCR. This study was funded by the Research
University Grant Scheme (UPM) [04-02-12-1796RU] and the Exploratory Research
Grant Scheme (Ministry of Higher Education Malaysia) [ERGS/1/2012/5527106].
Randy Haw Tatt Yhew, Tong Chih Kong an d Lee Ha n Chung are supp orted
by MyBr ain15 postgraduate scholarship programmes by the Ministry of
Education (MOE), Malaysia.
Author details
1
Neuroinflammation Group, Immunology Laboratory, Department of
Pathology, Faculty of Medicine & Health Sciences, Universiti Putra Malaysia,
43400 Serdang, Selangor, Malaysia.
2
Genetic & Regenerative Medicine
Research Centre (GRMRC) & Department of Obstetrics & Gynaecology,
Faculty of Medicine & Health Sciences, Universiti Putra Malaysia, 43400 UPM
Serdang, Selangor, Malaysia.
3
Department of Biomaterials & Tissue
Engineering, University College London, UCL Eastman Dental Institute,
256 Gray’s Inn Road, London WC1X 8LD, UK.
Received: 16 December 2013 Accepted: 16 July 2014
Published: 30 July 2014
References
1. Ginhoux F, Greter M, Leboeuf M, Nandi S, See P, Gokhan S, Mehler MF,
Conway SJ, Ng LG, Stanley ER, Samokhvalov IM, Merad M: Fate mapping
analysis reveals that adult microglia derive from primitive macrophages.
Science 2010, 330:841–845.
2. Kierdorf K, Erny D, Goldmann T, Sander V, Schulz C, Perdiguero EG,
Wieghofer P, Heinrich A, Riemke P, Holscher C, Muller DN, Luckow B,
Brocker T, Debowski K, Fritz G, Opdenakker G, Diefenbach A, Biber K,
Heikenwalder M, Geissmann F, Rosenbauer F, Prinz M: Microglia emerge
from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways.
Nat Neurosci 2013, 16:273–280.
3. Ransohoff RM, Cardona AE: The myeloid cells of the central nervous
system parenchyma. Nature 2010, 468:253–262.
4. Nimmerjahn A, Kirchhoff F, Helmchen F: Resting microglial cells are highly
dynamic surveillants of brain parenchyma in vivo. Science 2005,
308:1314–1318.
5. Kettenmann H, Hanisch UK, Noda M, Verkhratsky A: Physiology of
microglia. Physiol Rev 2011, 91:461–553.
6. Swinnen N, Smolders S, Avila A, Notelaers K, Paesen R, Ameloot M, Brone B,
Legendre P, Rigo JM: Complex invasion pattern of the cerebral cortex
bymicroglial cells during development of the mouse embryo. Glia 2013,
61:150–163.
7. Paolicelli RC, Bolasco G, Pagani F, Maggi L, Scianni M, Panzanelli P, Giustetto
M, Ferreira TA, Guiducci E, Dumas L, Ragozzino D, Gross CT: Synaptic
pruning by microglia is necessary for normal brain development. Science
2011, 333:1456–1458.
8. Schafer DP, Lehrman EK, Kautzman AG, Koyama R, Mardinly AR, Yamasaki R,
Ransohoff RM, Greenberg ME, Barres BA, Stevens B: Microglia sculpt
postnatal neural circuits in an activity and complement-dependent
manner. Neuron 2012, 74:691–705.
9. Sierra A, Encinas JM, Deudero JJ, Chancey JH, Enikolopov G,
Overstreet-Wadiche LS, Tsirka SE, Maletic-Savatic M: Microglia shape adult
hippocampal neurogenesis through apoptosis-coupled phagocytosis.
Cell Stem Cell 2010, 7:483–495.
10. Chuang HN, Van Rossum D, Sieger D, Siam L, Klemm F, Bleckmann A,
Bayerlova M, Farhat K, Scheffel J, Schulz M, Dehghani F, Stadelmann C,
Hanisch UK, Binder C, Pukrop T: Carcinoma cells misuse the host tissue
damage response to invade the brain. Glia 2013, 61:1331–1346.
11. Krabbe G, Halle A, Matyash V, Rinnenthal JL, Eom GD, Bernhardt U, Miller KR,
Prokop S, Kettenmann H, Heppner FL: Functional impairment of microglia
coincides with Beta-amyloid deposition in mice with Alzheimer-like
pathology. PLoS One 2013, 8:e60921.
12. Brown RA: In the beginning there were soft collagen-cell gels:
towards better 3D connective tissue models? Exp Cell Res 2013,
319:2460–24 69.
13. East E, De Oliveira DB, Golding JP, Phillips JB: Alignment of astrocytes
increases neuronal growth in three-dimensional collagen gels and is
maintained following plastic compression to form a spinal cord repair
conduit. Tissue Eng Part A 2010, 16:3173–3184.
14. East E, Golding JP, Phillips JB: A versatile 3D culture model facilitates
monitoring of astrocytes undergoing reactive gliosis. J Tissue Eng Regen
Med 2009, 3:634–646.
15. Phillips JB, Brown R: Structured Materials and Mechanical Cues in 3D
Collagen Gels. In 3D Cell Culture: Methods and Protocols, Methods in
Molecular Biology. Edited by Haycock JW. New York Dordrecht Heidelberg
London: Springer Science and Business Media; 2011:183–196.
16. Blasi E, Barluzzi R, Bocchini V, Mazzolla R, Bistoni F: Immortalization of murine
microglial cells by a v-raf/v-myc carrying retrovirus. JNeuroimmunol1990,
27:229–237.
17. Lizarraga F, Poincloux R, Romao M, Montagnac G, Le Dez G, Bonne I, Rigaill
G, Raposo G, Chavrier P: Diaphanous-related formins are required for
invadopodia formation and invasion of breast tumor cells. Cancer Res
2009, 69:2792–2800.
18. Hewitt CA, Ling KH, Merson TD, Simpson KM, Ritchie ME, King SL, Pritchard
MA, Smyth GK, Thomas T, Scott HS, Voss AK: Gene network disruptions
and neurogenesis defects in the adult Ts1Cje mouse model of Down
syndrome. PLoS One 2010, 5:e11561.
19. Olson JK, Miller SD: Microglia initiate central nervous system innate and
adaptive immune responses through multiple TLRs. J Immunol 2004,
173:3916–3924.
20. Dheen ST, Jun Y, Yan Z, Tay SS, Ling EA: Retinoic acid inhibits
expression of TNF-alpha and iNOS in activated rat microglia.
Glia 2005, 50:21–31.
21. Kim J, Hematti P: Mesenchymal stem cell-educated macrophages: a novel
type of alternatively activated macrophages. Exp Hematol 2009,
37:1445–1453.
22. Loane DJ, Stoica BA, Pajoohesh-Ganji A, Byrnes KR, Faden AI: Activation of
metabotropic glutamate receptor 5 (mGLUR5) modulates microglial
reactivity and neurotoxicity by inhibiting NADPH oxidase. J Biol Chem
2009, 284:15629–15639.
23. Ooi YY, Ramasamy R, Rahmat Z, Subramaiam H, Tan SW, Abdullah M,
Israf DA, Vidyadaran S: Bone marrow-derived mesenchymal stem cells
modulate BV2 microglia responses to lipopolysaccharide.
Int Immunopharmacol 2010, 10:1532–1540.
24. Rahmat Z, Jose S, Ramasamy R, Vidyadaran S: Reciprocal interactions of
mouse bone marrow-derived mesenchymal stem cells and BV2 microglia
after lipopolysaccharide stimulation. Stem Cell Res Ther 2013, 4:12.
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 9 of 10
http://www.jneuroinflammation.com/content/11/1/134

25. Vidyadaran S, Ooi YY, Subramaiam H, Badiei A, Abdullah M, Ramasamy R,
Seow HF: Effects of macrophage colony-stimulating factor on microglial
responses to lipopolysaccharide and beta amyloid. Cell Immunol 2009,
259:105–110.
26. Brown GC, Bal-Price A: Inflammatory neurodegeneration mediated by
nitric oxide, glutamate, and mitochondria. Mol Neurobiol 2003,
27:325–355.
doi:10.1186/1742-2094-11-134
Cite this article as: Haw et al.: A three-dimensional collagen construct to
model lipopolysaccharide-induced activation of BV2 microglia. Journal of
Neuroinflammation 2014 11:134.
Submit your next manuscript to BioMed Central
and take full advantage of:
• Convenient online submission
• Thorough peer review
• No space constraints or color figure charges
• Immediate publication on acceptance
• Inclusion in PubMed, CAS, Scopus and Google Scholar
• Research which is freely available for redistribution
Submit your manuscript at
www.biomedcentral.com/submit
Haw et al. Journal of Neuroinflammation 2014, 11:134 Page 10 of 10
http://www.jneuroinflammation.com/content/11/1/134
